- Research
- Open access
- Published:
Development and application of perfect SSR markers in cotton
Journal of Cotton Research volume 3, Article number: 21 (2020)
Abstract
Background
This study aimed to develop a set of perfect simple sequence repeat (SSR) markers with a single copy in the cotton genome, to construct a DNA fingerprint database suitable for authentication of cotton cultivars. We optimized the polymerase chain reaction (PCR) system for multi-platform compatibility and improving detection efficiency. Based on the reference genome of upland cotton and 10× resequencing data of 48 basic cotton germplasm lines, single-copy polymorphic SSR sites were identified and developed as diploidization SSR markers. The SSR markers were detected by denaturing polyacrylamide gel electrophoresis (PAGE) for initial screening, then fluorescence capillary electrophoresis for secondary screening. The final perfect SSR markers were evaluated and verified using 210 lines from different sources among Chinese cotton regional trials.
Results
Using bioinformatics techniques, 1 246 SSR markers were designed from 26 626 single-copy SSR loci. Adopting a stepwise (primary and secondary) screening strategy, a set of 60 perfect SSR markers was selected with high amplification efficiency and stability, easy interpretation of peak type, multiple allelic variations, high polymorphism information content (PIC) value, uniform chromosome distribution, and single-copy characteristics. A multiplex PCR system was established with ten SSR markers using capillary electrophoresis detection.
Conclusions
A set of perfect SSR markers of cotton was developed and a high-throughput SSR marker detection system was established. This study lays a foundation for large-scale and standardized construction of a cotton DNA fingerprint database for authentication of cotton varieties.
Background
Cotton is one of the most important economic crops in the world. Its fiber is a renewable textile raw material, and its seeds can be used for the production and processing of cottonseed oil. In recent years, excellent new varieties have been developed and promoted, and the seed market has flourished. At the same time, brand infringement and the production and sale of fake and inferior seeds have continued to occur, and this has severely dampened the enthusiasm of innovators, increased the risks for farmers, and affected agricultural production safety, which has severely restricted the development of China’s seed industry (Wu et al. 2015). Variety identification can effectively ensure the quality of seeds in agricultural production and reduce the risk of using fake and/or inferior seeds. Among DNA molecular fingerprinting methods, simple sequence repeat (SSR) molecular marker technology is widely used in rice (Sundaram et al. 2008; Singh et al. 2004), wheat (Zhu and Jia 2003; Shi et al. 2006), maize (Wang et al. 2003, 2017), soybean (Guan et al. 2003; Sheng et al. 2010), rape (Li et al. 2010) and other major crops because of its advantages such as high polymorphism, codominant inheritance, and easy statistics (Guichoux et al. 2011). There have numerous reports on SSR molecular detection technology in cotton (Pan et al. 2008; Kuang et al. 2011a, b; Wang 2009; Wang et al. 2009; Bai et al. 2012; Liu et al. 2009). However, in previous studies, most of the SSR primers were from public databases. The quality of primers was uneven, and the number of screening and evaluation materials tested was relatively small. Using the existing methods, it is difficult to meet the needs of identification, standardization, and large-scale detection of cotton varieties currently planted in China.
Wang et al. first proposed the concept of excellent SSR markers in maize and provided several important evaluation indicators: (1) based on the development of resequencing data, the primer flanking sequence should be highly conserved; (2) suitable markers can effectively remove abnormal peak shapes such as N + 1 peaks and continuous peaks; (3) multiplex polymerase chain reaction (PCR) can be combined to improve detection efficiency; (4) data integration should be available using different platforms (Wang et al. 2016). For cotton cultivars, the complexity of the polyploid genome (Yang 2001; Wolfe 2001) and the reproductive characteristics of cross-pollination pose difficulties for the development of high-quality markers and accurate genotyping among different varieties.
In recent years, with the completion of the cotton reference genome sequence (Wang et al. 2012; Li et al. 2014; Zhang et al. 2015), the foundation has been laid for the development and application of high-quality markers for the whole genome. In this study, we used 10× resequencing data from 48 upland cotton genotypes. Concerning the evaluation index of excellent SSR markers in maize (Wang et al. 2016), combined with the polyploid characteristics of cotton, a batch of single-copy SSR markers was developed. Denaturing polyacrylamide gel electrophoresis and fluorescent capillary electrophoresis were used for primary screening and re-screening of primers, respectively. We successfully developed a set of single-copy excellent SSR primers that combines high efficiency and stability, easy-to-read peak shapes, multiple allelic variations, high polymorphism information content (PIC) values, and uniform chromosome distribution. Combining multiplex PCR amplification technology and capillary electrophoresis detection enabled a ten-fold capillary electrophoresis procedure. In this way, a high-throughput SSR marker detection system was established, laying the foundation for constructing a large-scale database of DNA fingerprints for cotton variety identification and authentication.
Methods
Experimental materials
This study used 689 cotton cultivars, including 395 core germplasm lines (Ma et al. 2018), 210 lines from national trials in different regions, 48 basic germplasm lines, and 12 sets of triplet materials comprising male parent, female parent, and F1 generation (Table 1).
SSR primers
The common primers used in this study were synthesized by Dalian Bao Biological Company, China, and the fluorescent primers were synthesized by Applied Biosystems, USA. For each pair of fluorescent primers, one of them was selected to be labeled with a fluorescent dye PET, NED, VIC, or FAM at the 5′ end.
DNA extraction
Genomic DNA was extracted from dry seeds of each cultivar using the method described by Kuang et al. (2010).
SSR amplification
Amplification system: 20 μL reaction volume comprised 10 μL of 2× Taq Plus Master Mix, 0.4 μmol·L− 1 SSR primer, and 80 ng DNA template.
Amplification procedure: pre-denaturation at 94 °C for 5 min; denaturation at 94 °C for 45 s, annealing at 60 °C for 45 s, extension at 72 °C for 45 s, for a total of 32 cycles; extension at 72 °C for 7 min.
Denaturing polyacrylamide gel electrophoresis (PAGE)
Denatured PCR products were separated on a 6% polyacrylamide gel and electrophoresed at 90 W for 1 h and then stained with silver nitrate (Kuang et al. 2011b).
Fluorescence capillary electrophoresis
A mixture of 1 μL PCR amplification product, 8.5 μL deionized formamide and 0.5 μL Liz-500 molecular standard was denatured at 95 °C for 5 min, cooled at 4 °C for 10 min, and then separated using an ABI 3130XL gene analyzer. The fluorescent capillary electrophoresis detection procedure was pre-electrophoresis at 15 kV for 3 min, 2 kV injection for 2 s, electrophoresis at 15 kV for 20 min.
Statistical analysis
The numbers of alleles, genetic diversity, and polymorphism information content (PIC) of SSR molecular markers were analyzed using PowerMarker 3.25 software. The discrimination ability of SSR primers is expressed by the PIC value: the larger the PIC value, the stronger the discrimination ability. The formula is: PIC = 1 − ΣPi2, where Pi refers to the frequency of the i-th allelic variation. Cluster analysis was performed on the similarity coefficient matrix and genetic distance matrix using the unweighted pair-group method with arithmetic means (UPGMA).
Results
Screening of single-copy SSR
Bioinformatics methods were used to evaluate 10× resequencing data of 48 cotton genotypes, and 26 266 single-copy SSR loci were obtained. Of these, 5 712 loci with conserved flanking sequences were suitable for designing SSR primers. Analyzing the number of alleles of each SSR primer sequence in 48 cotton germplasm lines, we found that the more alleles, the stronger ability of the primers to identify different varieties. At the same time, the smaller amplified product, the higher efficiency of PCR amplification, which is more beneficial for constructing a multiplex PCR system. Considering the above conditions and the chromosome distribution of the SSR primers, 1 246 pairs were selected from the 5 712 loci for further screening.
Preliminary screening by denaturing PAGE platform
For preliminary screening, 48 germplasm lines from three cotton-growing regions were used to screen 1 246 pairs of primers by denaturing polyacrylamide gel electrophoresis. A total of 398 polymorphic primers were screened and were distributed across the 26 chromosomes. Among them, 145 primers had high amplification efficiency, good reproducibility, clear main bands, and had no non-specific amplification, and single bands when tested in known cultivars. Some example electrophoresis bands are shown in Fig. 1.
DNA fragments amplified by selected SSR primers. Electrophoresis results obtained for 48 germplasm lines from three cotton-growing regions. Denatured PCR products were separated on a 6% polyacrylamide gel and then stained with silver nitrate. a primer PC44; b primer PC37; c primer PC25; d primer PC49
Rescreening by fluorescence capillary electrophoresis
For secondary screening, 395 cotton core germplasm lines (Ma et al. 2018) representing a broad range of genotypes were used to analyze the allelic variation of 145 pairs of candidate SSR primers (selected from the preliminary screening) using fluorescence capillary electrophoresis as the detection platform. Twelve sets of triplets comprising male parent, female parent, and F1 generation were used as materials to analyze the heterozygosity of the 145 pairs of candidate SSR primers in the hybrids. From the results of capillary electrophoresis, we selected primers with sharp single peaks, no interference peaks, no continuous peaks, and stable and easy-to-read peak shapes. Combining the above indicators, 60 pairs of excellent SSR primers with high allelic variation, strong discrimination ability, uniform chromosome distribution, high heterozygosity in hybrids, and easy-to-read peak shape were selected.
Polymorphism analysis of SSR markers
The selected 60 excellent SSR primers were all diploid single-copy primers and were evenly distributed on the 26 chromosomes of cotton A and D genomes. The primer amplification was efficient and stable. In the capillary electrophoresis detection platform, 88.3% gave sharp single peaks with easy-to-read peak shape. There are many allelic variations among the primers, the 60 pairs of excellent SSR primers contained 247 single-copy polymorphic alleles among 431 cotton lines. Each primer contained an average of 4.12 allelic variations, with a range of 2–13. Among them, primer PC07 had the most allelic variations, with 13 mutations, indicating that the site has a high frequency of mutation and rich genetic diversity. The average PIC value was 0.48, with a range of 0.2–0.8. The results of 12 sets of triplets showed that there were 24 pairs of primers with heterozygosity greater than 50%; among them, primer PC06 had the highest heterozygosity at 75%. Primer information is shown in Table 2.
Construction of a multiplex PCR system and ten-fold capillary electrophoresis
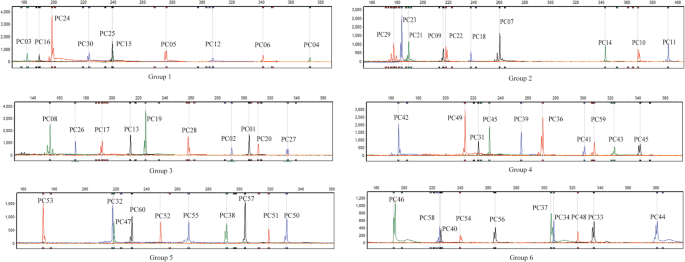
Primer Premier 5.0 software was used to analyze the interactions between the primers. Concerning the range of amplified fragments, 60 pairs of primers were divided into 18 combinations for multiplex PCR amplification. According to the different fluorescent colors of the primers, six groups of ten-fold primer combinations were constructed for capillary electrophoresis detection (Table 3).
Taking cotton variety Zhongmiansuo 49 as an example, the results of six ten-fold capillary electrophoresis detection are shown in Fig. 2. Ten pairs of SSR primers in each combination were labeled with four types of fluorescence. All the primers showed stable amplification, no interference between primers, clear bands and sharp peaks, making the results easy to interpret.
Application of perfect SSR markers in the construction of a DNA fingerprint database of cotton varieties
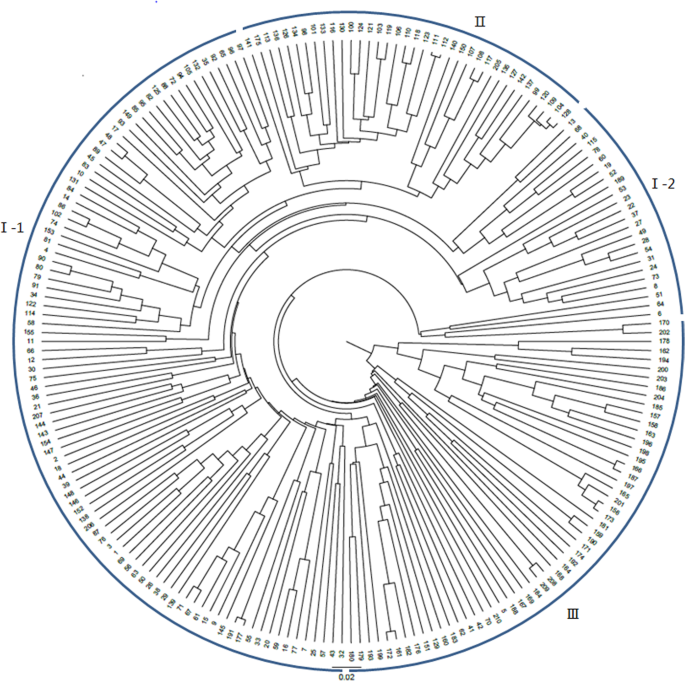
The selected 60 pairs of primers and the multiple combination method described above were used to construct fingerprints of 210 lines from national trials in different regions from 2016 to 2018. PowerMaker 3.25 software was used to calculate the genetic distance using Nei’s 1973 algorithm to construct a UPGMA clustering map. As shown in Fig. 3, samples No. 1–97 came from the Yellow River Basin; samples No. 98–146 came from the Yangtze River Basin; samples No. 147–210 came from the northwest inland region. The cluster analysis showed that: (1) the perfect SSR primer combinations were able to identify all 210 regional trial varieties; (2) regional trial samples from the same cotton-growing area were preferentially grouped.
Discussion
In recent years, molecular marker detection technologies have been continuously developed. Among such technologies, SSR and SNP molecular markers were written into the molecular testing guidelines by the International Union for the Protection of New Varieties of Plants (UPOV) in 2005 (UPOV 2005). At the same time, relatively mature SSR molecular markers have become the first choice for constructing DNA fingerprint databases of crops. The diploid crop maize is in a leading position in the construction of a standard fingerprint database of varieties using SSR molecular markers. The SSR fingerprint database and management system for certification of maize varieties have been constructed and applied to the national seed inspection system in China (Wang et al. 2017).
The selection of candidate markers and screening materials
In the case of cotton, the lack of genomic information in the early stage led to difficulty in the development of SSR primers, and the evaluation indexes for screening markers were insufficient. In the past, cotton SSR marker detection was mainly based on polyacrylamide gel electrophoresis, and there was no standardized operating procedure, which made it difficult to store and analyze the marker detection data efficiently, and the consistency of different laboratories was poor. Unlike diploid crops such as maize and rice, cultivated cotton is mostlytetraploid, so a single SSR marker may amplify into a single band, or two, three, or even four bands in different cotton varieties. At the same time, the breeding characteristic of frequent cross-pollination makes it difficult for cotton varieties to achieve high homozygosity, which further increases the difficulty of identifying cotton varieties using SSR molecular markers. In this study, we used 10× resequencing data of 48 basic cotton germplasm lines to develop SSR markers with single-copy characteristics and easy-to-read peak shapes, which amplify only a single or two bands in hybrids (characteristics similar to diploid crops), thereby greatly reducing the difficulty of data analysis and statistics.
The key factor in building a DNA fingerprint library of cotton varieties using SSR markers is the screening of a set of excellent SSR primers, which is largely dependent on the screening material. The origin of hetero-source tetraploid cultivated cotton varieties in China is mainly foreign seed, and the current production and breeding varieties are mainly derived from the USA, namely King cotton, Stoneville®, and Deltapine™ cotton varieties; the genetic basis is relatively narrow, with low genetic polymorphism between varieties (Guo et al. 2014). To obtain polymorphism information and accurate PIC values for the primers, this study took full account of a broad and representative source of genetic materials. We used 48 basic cotton lines from the Yellow River Basin, Yangtze River basin, and the northwest interior as the primary screening material, as well as nearly 400 core cotton germplasm lines representing 7 362 types of upland cotton (Ma et al. 2018) and 12 sets of triplets as re-screening materials. The final set of 60 pairs of excellent SSR primer had 247 single-copy isometric variant types, high PIC values, easy-to-interpret peak shapes, even chromosome distribution.
The specialties and application of perfect SSR markers
The selected 60 pairs of excellent SSR markers are compatible with both conventional polyacrylamide gel electrophoresis and fluorescent capillary electrophoresis detection platforms, and so are suitable for use by a wide range of researchers in different laboratory conditions. Compared with polyacrylamide gel electrophoresis detection, fluorescent capillary electrophoresis has the following advantages: (1) higher resolution, as fine as 1 bp; (2) electrophoresis results are displayed in a peak plot corresponding to the size of the amplified fragments, which can be directly collected to achieve digitization of the results in a form, that is easy to record, analyze and save; (3) the application of high-throughput, multi-PCR and 10-fold capillary electrophoresis detection technology has improved the detection efficiency by 33-fold. The 60 selected primers were used to establish a DNA fingerprinting library of 96 samples, which reduced the number of 96-well PCR plates needed for amplification from the original 60 to 18. Similarly, the number of 96-well electrophoresis plates required was reduced from the original 60 to 10, which greatly reduces the workload and testing costs; (4) the data are easy to integrate and are suitable for large-scale construction of a DNA fingerprint library.
We built a DNA fingerprint database of 210 samples from the Yellow River Basin, Yangtze River Basin and northwest inland cotton-growing regions using 60 markers. The results of genetic diversity analysis showed that the varieties in the same cotton region were mostly gathered together in the same group, but some varieties of the northwest inland region were scattered in group I, which mainly comprised varieties from the Yellow River basin. This might be because the ecological conditions of some parts of southern Xinjiang are similar with those of the Yellow River basin, or the existence of introduction exchange, which is similar to the results of Guo Zhijun’s study (Guo et al. 2014). In terms of identification capacity, using 60 pairs of excellent SSR primers enabled this method to distinguish 210 cotton national trial samples with a 100% success rate.
Conclusions
This study has provided a set of perfect SSR primers as well as optimized the PCR and electrophoresis system, which improved the detection efficiency by 33 times. It has laid an important foundation for the construction and application of a standardized cotton variety identification system on a large scale.
Availability of data and materials
The data and materials for supporting the results of this article are included within the article.
References
Bai J, Nie Y, Lin Z, et al. Selection and evaluation of SSR core primers in cotton hybrid. Cotton Sci. 2012;24(3):207–14. https://doi.org/10.3969/j.issn.1002-7807.2012.03.003.
Guan R, Liu Y, Liu Z, et al. Purity identification of soybean varieties with SSR technique. Mol Plant Breed. 2003;1(3):357–60. https://doi.org/10.3969/j.issn.1672-416X.2003.03.010.
Guichoux E, Lagache L, Wagner S, et al. Current trends in microsatellite genotyping. Mol Ecol Resour. 2011;11(4):591–611. https://doi.org/10.1111/j.1755-0998.2011.03014.x.
Guo Z, Zhao Y, Chen W, et al. Genetic diversity of SSR markers in upland cotton and its correlation with agronomic traits. Cotton Sci. 2014;26(5):420–30. https://doi.org/10.3969/j.issn.1002-7807.2014.05.006.
Kuang M, Yang W, Xu H, et al. Rapid extraction of DNA from single cotton seed. Mol Plant Breed. 2010;8(4):1–5. https://doi.org/10.3969/mpb.008.000827.
Kuang M, Yang W, Xu H, et al. Construction of DNA fingerprinting and analysis of genetic diversity with SSR markers for cotton major cultivars in China. Sci Agric Sin. 2011b;44(1):20–7. https://doi.org/10.3864/j.issn.0578-1752.2011.01.003.
Kuang M, Yang W, Zhang Y, et al. SSR marker detection of cotton hybrid purity and its correlation with field phenotype identification. Crop J. 2011a;37(12):2299–305. https://doi.org/10.3724/SP.J.1006.2011.02299.
Li F, Fan G, Wang K, et al. Genome sequence of the cultivated cotton Gossypium arboreum. Nat Genet. 2014;46(6):567–72. https://doi.org/10.1038/ng.2987.
Li H, Yang J, Lü Z, et al. Screening of Brassica napus core SSR primers. Chin J Oil Crop Sci. 2010;32(3):329–36.
Liu F, Feng X, Zhong W, et al. Screening of SSR core primer pairs for identificating cotton cultivar. Mol Plant Breed. 2009;7(6):1160–8. https://doi.org/10.3969/mpb.007.001160.
Ma Z, He S, Wang X, et al. Resequencing a core collection of upland cotton identifies genomic variation and loci influencing fiber quality and yield. Nat Genet. 2018;50:803–13. https://doi.org/10.1038/s41588-018-0119-7.
Pan Z, Sun J, Wang X, et al. Screening of SSR core primers with polymorphism on a cotton panel. Biodivers Sci. 2008;16(6):555–61. https://doi.org/10.3321/j.issn:1005-0094.2008.06.005.
Sheng X, Zhan S, Fan H, et al. Purity identification of “Meiqiao” soybean (Glycine max) by SSR markers. Soybean Sci. 2010;29(2):210–4. https://doi.org/10.11861/j.issn.1000-9841.2010.02.0210.
Shi H, Wang L, Li H, et al. Division of seeds motley and SSR loci impurity in wheat cultivar using SSR markers. Mol Plant Breed. 2006;4(4):513–9. https://doi.org/10.3969/j.issn.1672-416X.2006.04.011.
Singh RK, Sharma RK, Singh AK, et al. Suitability of mapped sequence tagged microsatellite site markers for establishing distinctness, uniformmity and stability in aromatic rice. Euphytica. 2004;135:135–43. https://doi.org/10.1023/B:EUPH.0000014905.10397.08.
Sundaram RM, Naveenkumar B, Biradar SK, et al. Identification of informative SSR markers capable of distinguishing hybrid rice parental lines and their utilization in seed purity assessment. Euphytica. 2008;163:215–24. https://doi.org/10.1007/s10681-007-9630-0.
UPOV (Union for the Protection of New Varieties of Plants). Guidelines for molecular marker selection and database construction, BMT Guidelines (Proj.3). Geneva: UPOV; 2005. p. 3–6.
Wang F, Yang Y, Yi H, et al. Construction of an SSR-based standard fingerprint database for corn variety authorized in China. Sci Agric Sin. 2017;50(1):1–14. https://doi.org/10.3864/j.issn.0578-1752.2017.01.001.
Wang F, Yi H, Zhao J, et al. Development and application of SSR identification standard for authenticity of corn. Corn Sci. 2016;24(4):61–6. https://doi.org/10.13597/j.cnki.maize.science.20160412.
Wang F, Zhao J, Guo J, et al. Comparison of three DNA fingerprint analyzing methods for maize cultivars identification. Mol Plant Breed. 2003;1(5):655–61. https://doi.org/10.3969/j.issn.1672-416X.2003.05.010.
Wang J. SSR markers-based cotton purity and authenticity identification. Beijing: Chinese Academy of Agricultural Sciences; 2009.
Wang J, Yang W, Kuang M, et al. Establishment of fingerprinting and research on identification for cotton varieties. China Cotton. 2009;36(3):6–9. https://doi.org/10.3969/j.issn.1000-632X.2009.03.002.
Wang K, Wang Z, Li F, et al. The draft genome of a diploid cotton Gossypium raimondii. Nat Genet. 2012;44(10):1098–103. https://doi.org/10.1038/ng.2371.
Wolfe K. Yesterday’s polyploids and the mystery of diploidization. Nat Rev Genet. 2001;2(5):333–41.
Wu X, Peng Z, He L. Thinking on solving the dilemma of cracking down on infringement and illegal acts from the legal level. China Seed Ind. 2015;9:1–3, 11. https://doi.org/10.19462/j.cnki.1671-895x.2015.09.001.
Yang J. The formation and evolution of polyploid genomes in plants. Acta Phytotax Sin. 2001;39(4):357–71.
Zhang T, Hu Y, Jiang W, et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fibre improvement. Nat Biotechnol. 2015;33(5):531–7. https://doi.org/10.1038/nbt.3207.
Zhu Z, Jia J. Development and application of wheat SSR markers. Hereditas. 2003;25(3):355–60. https://doi.org/10.16288/j.yczz.2003.03.026.
Acknowledgements
We would like to thank the anonymous reviewers for their valuable comments and helpful suggestions which help to improve the manuscript.
Funding
This work was supported by grants from the Thirteenth Five-Year Plan, National Key R & D Plan (2017YFD0102003–5); National Cotton Industry Technology System(CARS-15-25).
Author information
Authors and Affiliations
Contributions
Kuang M, Peng J initiated the research; Wu YZ and Huang LY designed the experiments; Wu YZ, Huang LY, Zhou DY, Fu XQ, Li C, and Wei SJ collected the data from the field; Wu YZ performed the analysis; Wu YZ drafted the manuscript; Kuang M and Peng J finalized the manuscript. All authors contributed in the interpretation of results and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
WU, Y., HUANG, L., ZHOU, D. et al. Development and application of perfect SSR markers in cotton. J Cotton Res 3, 21 (2020). https://doi.org/10.1186/s42397-020-00066-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s42397-020-00066-0